|
Size: 2031
Comment:
|
Size: 2617
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 21: | Line 21: |
* tip if you use Mozilla Firefox (e.g. 30.0) webbrowser: it can take more than 5 min to get the results back from the oPOSSUM server, in this case you need to change the default response timeout (which is 300s = 5 mins): * open Firefox * in the address bar, enter about:config and accept the warning message * set network.http.response.timeout to 0(no response time out or a time of your choice e.g 3000 for 3000 seconds) * reopen Firefox and launch oPOSSUM 3.0 |
|
| Line 34: | Line 41: |
| * [[CancerStemCellProject/VeroniqueVoisin/AdditionalResources/CHIP_seq | link to ChIP-seq basics]] |
Find over represented transcription factor motifs or binding sites in co-expressed genes
- Finding over-representation of transcription factor binding sites in group of genes/pathways found dysregulated in gene expression data
TOOLS THAT USE TRANSCRIPTION FACTOR MOTIFS
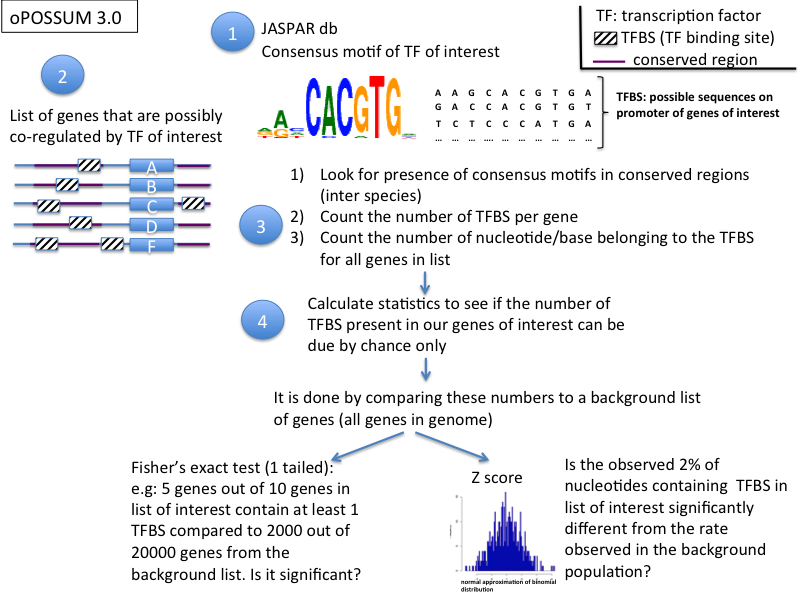
1) oPOSSUM (tested, 3.0) (http://www.cisreg.ca/oPOSSUM/)
- web tool (no account necessary)
- the method has 3 steps:
- phylogenetic footprinting to find regions in the non coding DNA (promoter regions) that are conserved between species
- detection of transcription factor motifs using the JASPAR database (JASPAR PSSMs: position specific scoring matrices)
- 2 statistics methods (Fisher's score and Z score) to evaluate over-represented binding sites compared to background
- tips:
- select potential transcription factor candidates by generating a Z score / Fisher's plot: select the transcription factors that emerged from the cloud
look at the %GC content - Z score to see if you have a %gC content bias: if any, run the GC_compo tool (http://opossum.cisreg.ca/GC_compo/) to select an appropriate background set and use the Sequence-based Single Site Analysis tool
- tip if you use Mozilla Firefox (e.g. 30.0) webbrowser: it can take more than 5 min to get the results back from the oPOSSUM server, in this case you need to change the default response timeout (which is 300s = 5 mins):
- open Firefox
- in the address bar, enter about:config and accept the warning message
- set network.http.response.timeout to 0(no response time out or a time of your choice e.g 3000 for 3000 seconds)
- reopen Firefox and launch oPOSSUM 3.0
2) PSCAN: http://159.149.109.9/pscan/
- need Refseq as gene identifier
TOOLS THAT USE ENCODE CHIP-seq DATA
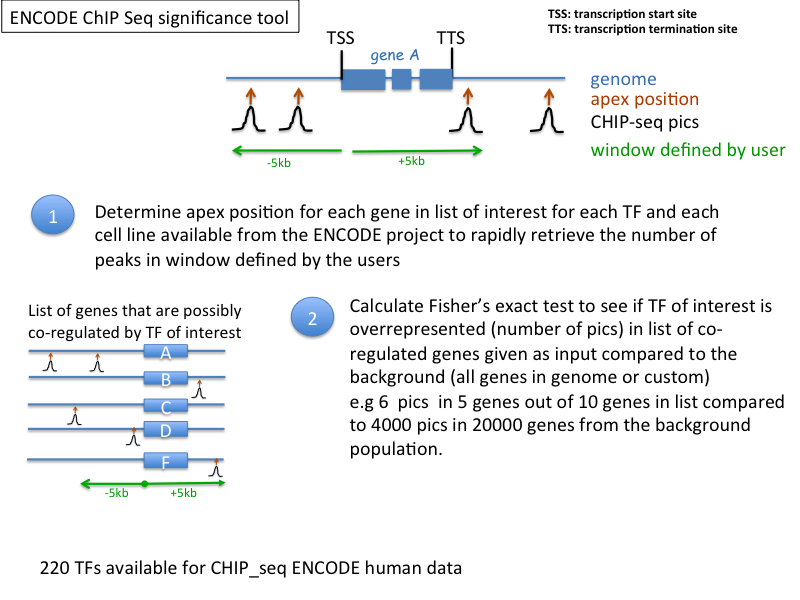
ENCODE ChIP-Seq Significance Tool: http://encodeqt.stanford.edu/hyper/
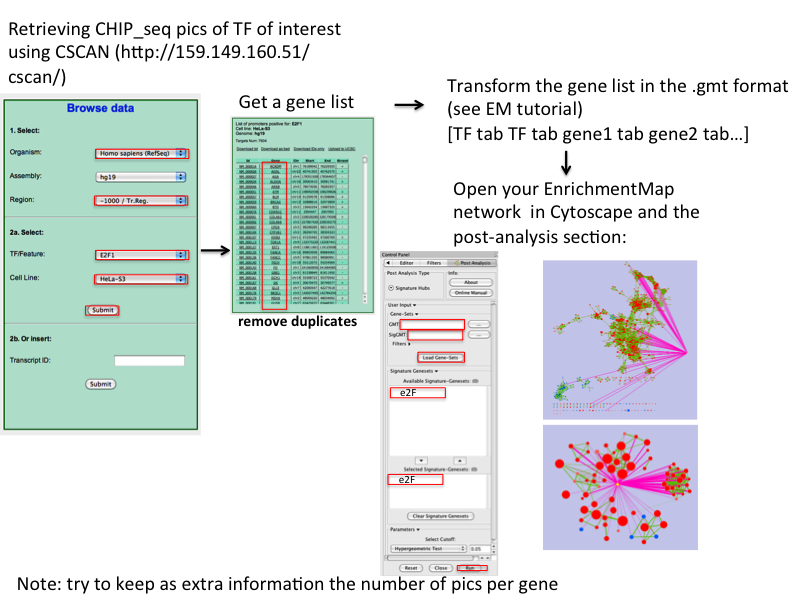
CSAN: (as PSCAn but using chip-seq data): http://159.149.109.9/cscan/
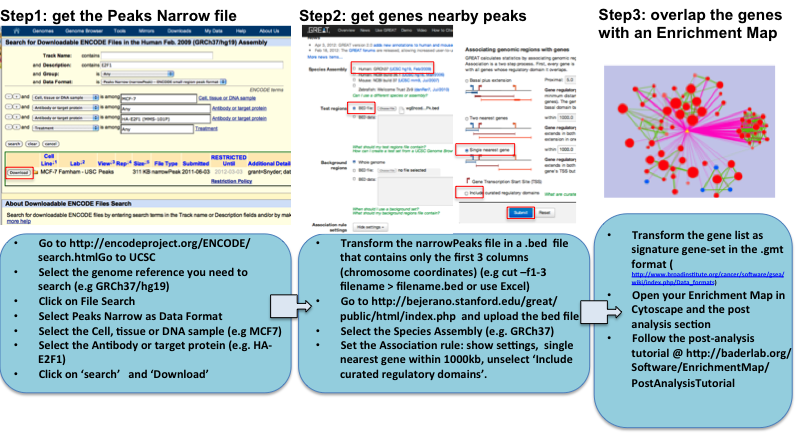
GET THE GENES NEARBY ChIP-SEQ ENCODE DATA and LOOK AT THE MAP OVERLAP using EnrichmentMap POST-ANALYSIS
REFERENCES:
