Find over represented transcription factor motifs in co-expressed genes
- Finding over-representation of transcription factor binding sites in group of genes/pathways found dysregulated in gene expression data
TOOLS THAT US TRANSCRIPTION FACTOR MOTIFS
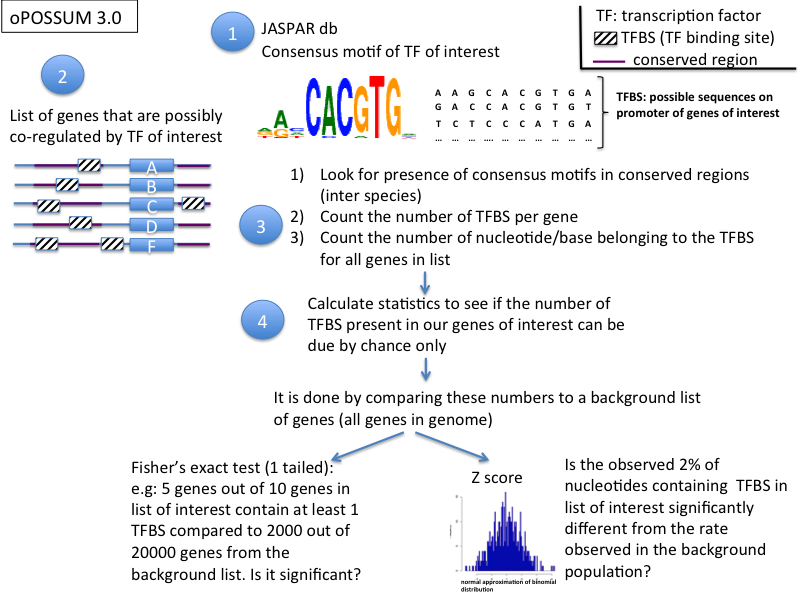
1) oPOSSUM (tested, 3.0) (http://www.cisreg.ca/oPOSSUM/)
- web tool (no account necessary)
- the method has 3 steps:
- phylogenetic footprinting to find regions in the non coding DNA (promoter regions) that are conserved between species
- detection of transcription factor motifs using the JASPAR database (JASPAR PSSMs: position specific scoring matrices)
- 2 statistics methods (Fisher's score and Z score) to evaluate over-represented binding sites compared to background
- tips:
- select potential transcription factor candidates by generating a Z score / Fisher's plot: select the transcription factors that emerged from the cloud
look at the %GC content - Z score to see if you have a %gC content bias: if any, run the GC_compo tool (http://opossum.cisreg.ca/GC_compo/) to select an appropriate background set and use the Sequence-based Single Site Analysis tool
2) PSCAN: http://159.149.109.9/pscan/
- need Refseq as gene identifier
TOOLS THAT USE ENCODE CHIP-seq DATA
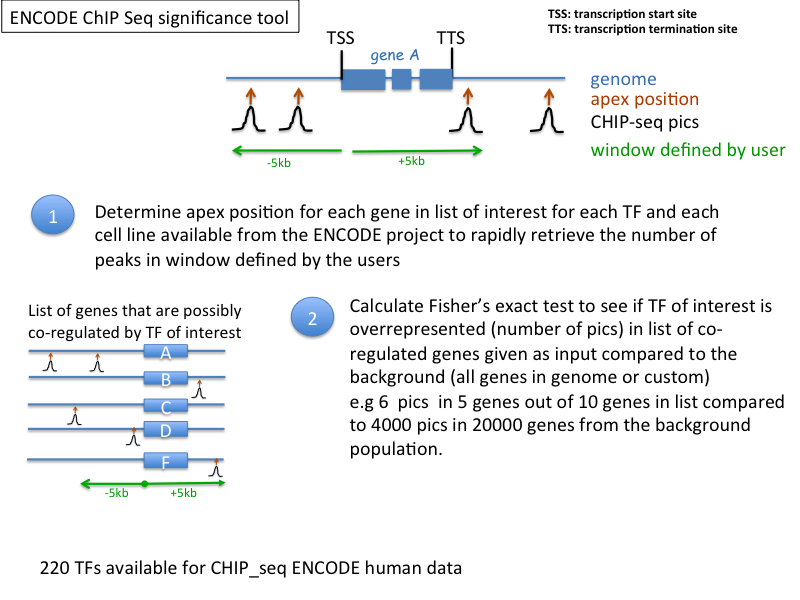
ENCODE ChIP-Seq Significance Tool: http://encodeqt.stanford.edu/hyper/
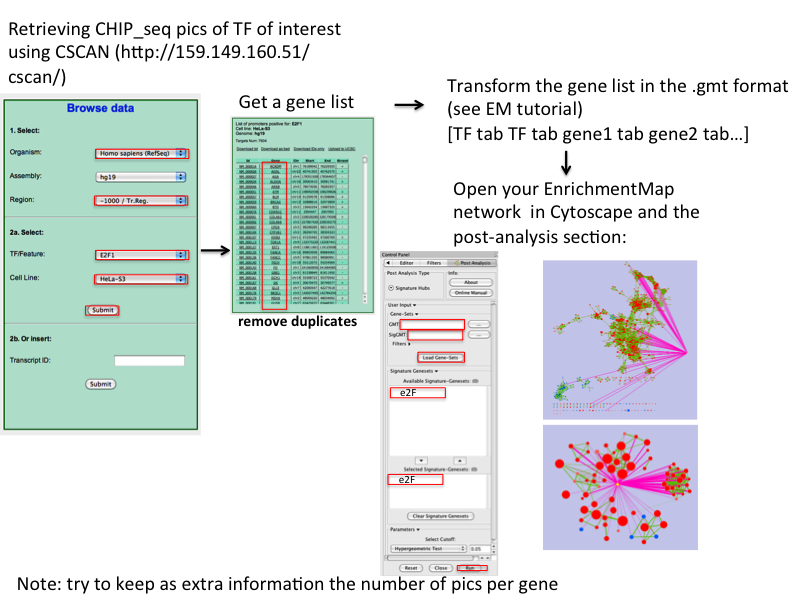
CSAN: (as PSCAn but using chip-seq data): http://159.149.109.9/cscan/
REFERENCES:
Wyeth Wasserman lab: http://www.cisreg.ca/
blog: http://gettinggeneticsdone.blogspot.ca/2013/06/encode-chip-seq-significance-tool-which.html