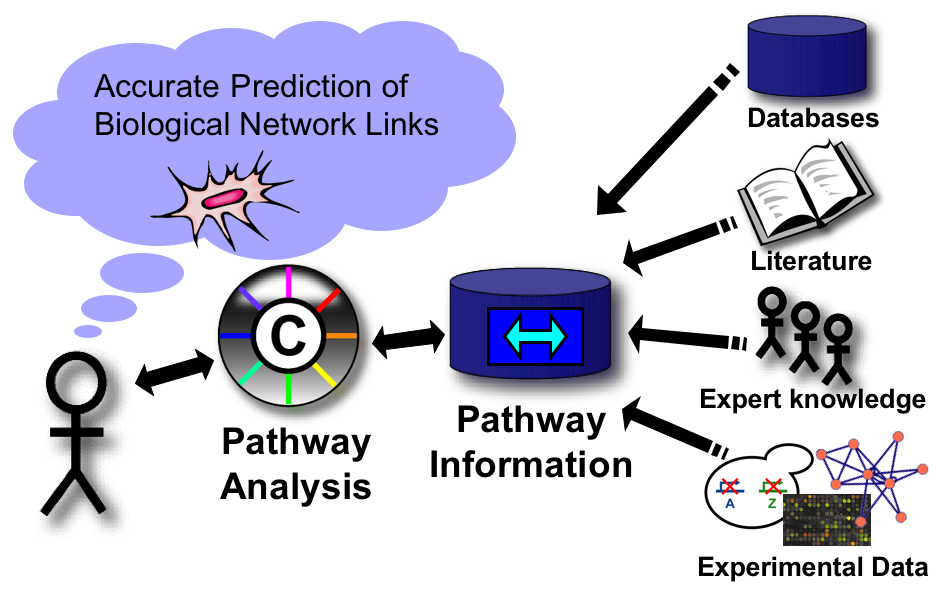
The Bader Lab is involved in a number of collaborative open-source bioinformatics projects designed to make biological pathway data easy to visualize and analyze.
Contents
Biological Network Analysis and Visualization Software
Cytoscape
Cytoscape is a bioinformatics software platform for visualizing molecular interaction networks and integrating these interactions with gene expression profiles and other state data. Additional features are available as plugins. Plugins are available for network and molecular profiling analyses, new layouts, additional file format support and connection with databases. Plugins may be developed using the Cytoscape open Java software architecture by anyone and plugin community development is encouraged.
Cytoscape was originally developed at the Institute of Systems Biology and is now a collaborative effort involving many different academic and commercial groups.
Fun link: Controlling Cytoscape using the Nintendo Wiimote
NetMatch
NetMatch is a Cytoscape plugin that finds user defined network motifs in any Cytoscape network. Node and edge attributes of any type and paths of unknown length can be specified in the search.
Released by: Ferro, Giugno, Pulvirenti group, University of Catania, Bader group, University of Toronto and Shasha group, New York University.
Available from the Bader Lab mirror of the NetMatch homepage
MCODE
MCODE is a Cytoscape plugin that finds clusters (highly interconnected regions) in a network. Clusters mean different things in different types of networks. For instance, clusters in a protein-protein interaction network are often protein complexes and parts of pathways, while clusters in a protein similarity network represent protein families.
Available from the MCODE homepage
BRAIN
The Biologically Relevant Analysis of Interaction Networks (BRAIN) is a set of algorithms for predicting and analyzing protein domain-peptide ligand interactions based on experimentally known binding evidence (e.g. from protein chip or phage display experiments). BRAIN can be accessed as a Cytoscape plugin which reads peptide binding profiles and generates interactions displayed as a Cytoscape network.
Available from the BRAIN homepage
LOLA
LOLA (LOgos Look Amazing) is a tool for generating sequence logos using Position Weight Matrix based protein profiles. LOLA allows you to generate custom sequence logos by setting parameters such as logo height, trim percentage, and residue colour scheme. You can also generate "logo trees" based on clustered protein profiles to analyze binding motif classes. Sequence logos and logo trees can be saved in various formats including PDF, PNG, and JPEG.
Available from the LOLA homepage
Enrichment Map
Enrichment Map is a Cytoscape plugin for functional enrichment visualization. Enrichment results have to be generated outside Enrichment Map, using any of the available methods. Gene-sets, such as pathways and Gene Ontology terms, are organized into a network (i.e. the "enrichment map"). In this way, mutually overlapping gene-sets cluster together, making interpretation easier. Enrichment Map also enables the comparison of two different enrichment results in the same map.
Available from Enrichment Map homepage
WordCloud
WordCloud is a Cytoscape app that generates a visual summary of a network. It displays string attributes associated with nodes in the network as a tag cloud, where more frequent words are displayed using a larger font size. Word co-occurence in a phrase can be visualized by arranging words in clusters or as a network.
Available from WordCloud homepage
AutoAnnotate
AutoAnnotate is a Cytoscape app that finds node clusters and visually annotates them with labels and groups. These visualizations provide a concise visual summary which is helpful for network analysis and interpretation.
Available from AutoAnnotate homepage
SIREN
The Signing of Regulatory Networks (SIREN) algorithm can infer the regulatory type (positive or negative regulation) of interactions in a known gene regulatory network given corresponding genome-wide gene expression data.
http://apps.cytoscape.org/apps/siren
SocialNetworkApp
The Social Network Cytoscape app creates a visual summary of how individuals are connected. Biological networks can be visualized and analyzed using Cytoscape. Often researchers want to go beyond the network of proteins or genes and also look at the inter-connectedness between colleagues and institutions. Who tends to publish together? What institutions are most collaborative? Are there inter-disciplinary connections in my institution? The app addresses these questions by building co-publication networks where the nodes represent authors, edges represent co-authorship and edge thickness represents how frequently co-authors collaborate.
Available from SocialNetworkApp homepage
Biological Pathway and Network Database Software
GeneMANIA
GeneMANIA helps you predict the function of your favourite genes and gene sets.
GeneMANIA finds other genes that are related to a set of input genes, using a very large set of functional association data. Association data include protein and genetic interactions, pathways, co-expression, co-localization and protein domain similarity. You can use GeneMANIA to find new members of a pathway or complex, find additional genes you may have missed in your screen or find new genes with a specific function, such as protein kinases. Your question is defined by the set of genes you input.
This work is done in collaboration with Quaid Morris' lab at the University of Toronto.
cPath
cPath, an open source database and web application for collecting, storing, browsing and querying biological pathway data. cPath makes it easy to aggregate custom pathway data sets available in standard exchange formats from multiple databases, present pathway data to biologists via a customizable web interface, and easily export pathway data via a web service to third-party software for visualization and analysis. cPath is software only, and does not include new pathway information. Key features include: a built-in identity and reference service for identifying and annotating interactors; built-in support for PSI-MI and BioPAX standard pathway exchange formats; and a web service interface for searching and retrieving pathway data sets. Thorough documentation is available for all aspects of cPath. The cPath software is freely available under the LGPL open source license for academic and commercial use.
http://code.google.com/p/pathway-commons/
http://www.pathwaycommons.org/ (Pathways and molecular interactions)
PC (cPath1): http://www.pathwaycommons.org/pc/
PC2 (cPath2): http://www.pathwaycommons.org/pc2/
This work is done in collaboration with the Sander group of the Computational Biology Center at Memorial Sloan-Kettering Cancer Center in New York City.
Pathway and Network Data
Pathway Commons
Pathway Commons is a convenient point of access to biological pathway information collected from public pathway databases, which you can browse or search. Pathways include biochemical reactions, complex assembly, transport and catalysis events, and physical interactions involving proteins, DNA, RNA, small molecules and complexes.
http://www.pathwaycommons.org/
This work is done in collaboration with the Sander group of the Computational Biology Center at Memorial Sloan-Kettering Cancer Center in New York City.
The Cancer Cell Map
The Cancer Cell Map contains selected cancer related signaling pathways which you can browse or search. Biologists can browse and search the Cancer Cell Map pathways. View gene expression data on any pathway. Computational biologists can download all pathways in BioPAX format for global analysis. Software developers can build software on top of the Cancer Cell Map using the web service API. Download and install the cPath software to create a local mirror.
This work is done in collaboration with the Sander group of the Computational Biology Center at Memorial Sloan-Kettering Cancer Center in New York City.
Pathway Data Exchange Standards
BioPAX
BioPAX (Biological Pathway Exchange) is a collaborative effort to create a data exchange format for biological pathway data. BioPAX covers metabolic pathways, molecular interactions and protein post-translational modifications. Future versions will expand support for signaling pathways, gene regulatory networks and genetic interactions.
PSI-MI
PSI-MI (The Proteomics Standards Initiative Molecular Interactions) format allows exchange of molecular interaction data, focusing on protein-protein interactions.
http://psidev.sourceforge.net/
Both of these projects are large international collaborative efforts involving many different academic and commercial groups.
Community Web Sites
Pathguide
Pathguide, the Pathway Resource List, contains information about hundreds of online biological pathway resources. Databases that are free and those supporting BioPAX, CellML, PSI-MI or SBML standards are highlighted.
Vision
The above software support a vision for easy, seamless access to all public pathway information for visualization and interpretation in biological research.