Enrichment Map Plugin Download Page
Contents
Brief Description
Enrichment Map is a Cytoscape plugin for functional enrichment visualization. Enrichment results have to be generated outside Enrichment Map, using any of the available methods. Gene-sets, such as pathways and Gene Ontology terms, are organized into a network (i.e. the "enrichment map"). In this way, mutually overlapping gene-sets cluster together, making interpretation easier. Enrichment Map also enables the comparison of two different enrichment results in the same map.
Follow this link for more details and an analysis example.
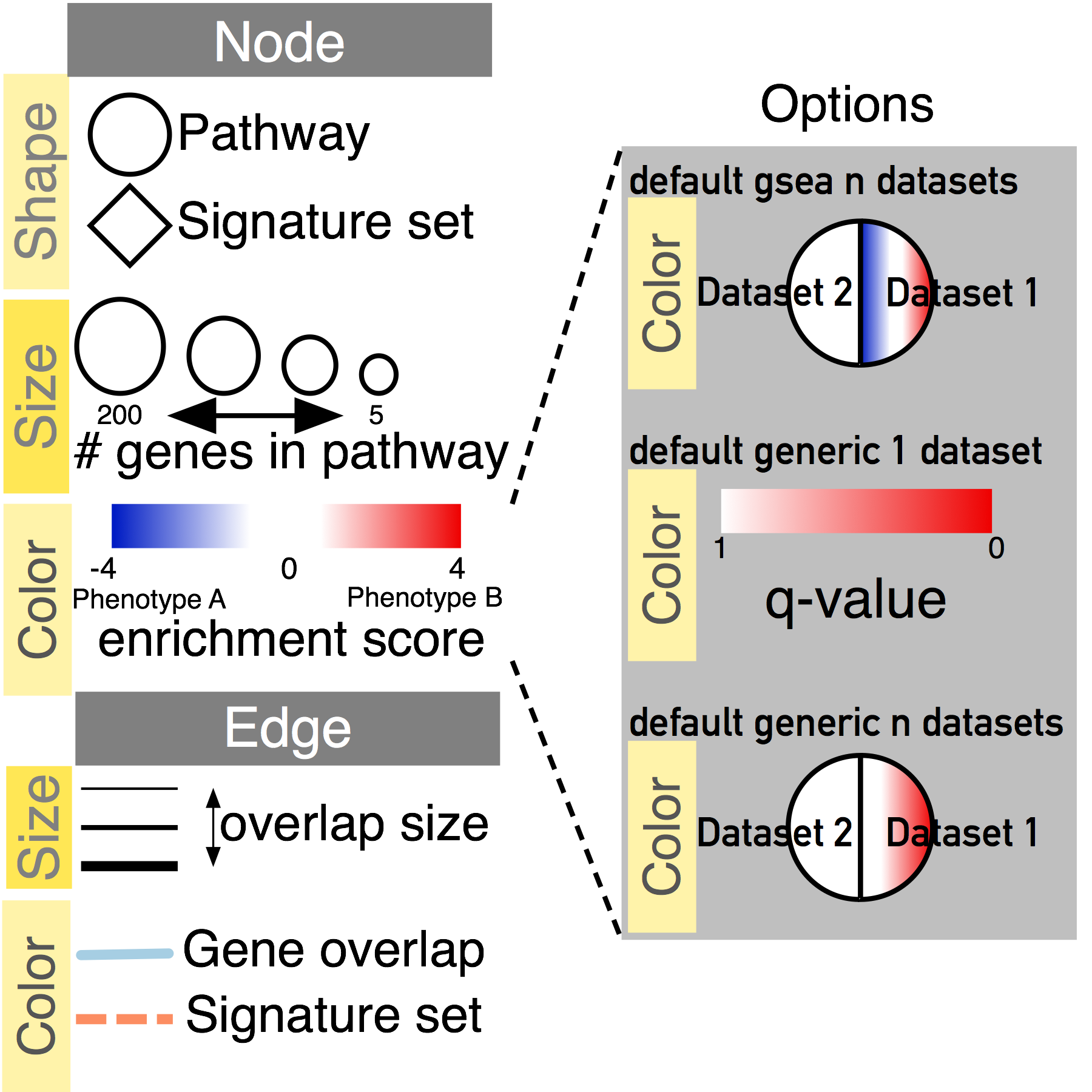
Legends
For easier figure creation download our template legends that contains all the visual properties used in a standard enrichment map. Download and customize legend to fit your network
Downloads
Cytoscape can be downloaded from: http://cytoscape.org/
EnrichmentMap is available on the Cytoscape App Store: http://apps.cytoscape.org/apps/enrichmentmap
Alternatively download the EnrichmentMap Pipeline Collection. This will install EnrichmentMap as well as a suite of other apps that work well with EnrichmentMap, including AutoAnnotate, WordCloud and clusterMaker2: http://apps.cytoscape.org/apps/enrichmentmappipelinecollection
Tutorials and Examples
Tuturials and protocols for EnrichmentMap 3.0 are here: https://baderlab.github.io/Cytoscape_workflows/EnrichmentMapPipeline/
Tutorials for the older 2.2 version of EnrichmentMap are here: http://enrichmentmap.readthedocs.io/en/docs-2.2/Tutorials.html
User Guide
The EnrichmentMap User Guide has been moved to: http://enrichmentmap.readthedocs.io
User Guide for older 2.2 version of EnrichmentMap (including tutorials) is here: http://enrichmentmap.readthedocs.io/en/docs-2.2/
Gene-sets for Enrichment Analysis
directly downloading our monthly updated gene-set collections from Baderlab genesets collections. Description of sources and methods used to create collection can be found here
- we generate gene-set files for enrichment analysis (e.g. GSEA) by querying public resources such as Gene Ontology and KEGG, and using Entrez-Gene ID for genes
how to convert DAVID gene-sets to GMT: R script
Publications
Cite EM
Enrichment Map: A Network-Based Method for Gene-Set Enrichment Visualization and Interpretation
Merico D, Isserlin R, Stueker O, Emili A, Bader GD
PLoS One. 2010 Nov 15;5(11):e13984
PubMed Abstract -PDF
Examples of Use
Functional impact of global rare copy number variation in autism spectrum disorders.
Pinto D, Pagnamenta AT, Klei L, Anney R, Merico D, Regan R, Conroy J, Magalhaes TR, Correia C, Abrahams BS, Almeida J, Bacchelli E, Bader GD, et al.
Nature. 2010 Jun 9 (Epub ahead of print)
PubMed Abstract - PDF
Nature Blogs - In the news (The Globe and Mail)
Pathway analysis of dilated cardiomyopathy using global proteomic profiling and enrichment maps
Isserlin R, Merico D, Alikhani-Koupaei R, Gramolini A, Bader GD, Emili A.
Proteomics 2010, March 10(6):1316-27
Pubmed Abstract - PDF
Papers Citing Enrichment Map
Pathway analysis of expression data: deciphering functional building blocks of complex diseases.
Emmert-Streib F, Glazko GV.
PLoS Comput Biol. 2011 May;7(5):e1002053.
PubMedInflammasome is a central player in the induction of obesity and insulin resistance.
Stienstra R, van Diepen JA, Tack CJ, Zaki MH, van de Veerdonk FL, Perera D, Neale GA, Hooiveld GJ, Hijmans A, Vroegrijk I, van den Berg S, Romijn J, Rensen PC, Joosten LA, Netea MG, Kanneganti TD.
Proc Natl Acad Sci U S A. 2011 Aug 29.
PubMedDelineation of Two Clinically and Molecularly Distinct Subgroups of Posterior Fossa Ependymoma
Witt H, Mack SC, Ryzhova M, Bender S, Sill M, Isserlin R, Benner A, Hielscher T, Milde T, Remke M, Jones DTW, Northcott PA, Garzia L, Bertrand KC, Wittmann A, Yao Y, Roberts SS, Massimi L, Van Meter T, Weiss WA, Gupta N, Grajkowska W, Lach B, Cho YJ, von Deimling A, Kulozik AE, Witt O, Bader GD, Hawkins CE, Tabori U, Guha A, Rutka JT, Lichter P, Korshunov A, Taylor MD, Pfister SM
Cancer Cell, Volume 20, Issue 2, 143-157, 16 August 2011
PubMed Abstract - PDF
Report a Bug or a Problem
please make sure you don't have formatting issues (have a look at the User Guide (and FAQ))
- if you are still not sure how to handle formats,
- or you don't know what's the best suitable analysis for you,
- please send an email to: daniele[AT]merico[DOT]gmail.com
- please check what's your
- plugin version and build (from the Cytoscape menu / Plugins / Enrichment Map / About)
- Cytoscape version (from the Cytoscape menu / Cytoscape)
- Operating System (e.g. Windows Vista)
and send your bug report to ruth[DOT]isserlin[AT]utoronto.ca
OR
report the bug to the Enrichment map issue tracker:- click on "Issues"
- click on "New Issue"
- write a short description of the issue
- attached session file (.cys) file or example input files if applicable
- Make sure to enter plugin version and build, cytoscape version and operating system.
- click on "Submit new issue"
For Feature requests please send to enrichmentmap-dev@googlegroups.com